Stanford Wants You to Play a Video Game to Improve CRISPR

Stanford needs you to play video games. Seriously. In order to improve CRISPR, the famous gene-editing tool, a research group at the Stanford University School of Medicine has created a new challenge for their online video game, EteRNA.
The end goal is an RNA molecule that can act as an on/off switch for CRISPR. Stanford molecular biologists will build the most promising player-submitted molecule designs and test them out. So if you love video games, what are you waiting for? The future of genetics needs you.

Gaming for Science
Stanford University researchers first launched EteRNA as an experiment a few years ago in 2011. The original intent of the game was not only to entertain but to educate people about genetics. The “stretch goal” was to have thousands of players designing enough new RNA sequences that the results would be clinically significant.
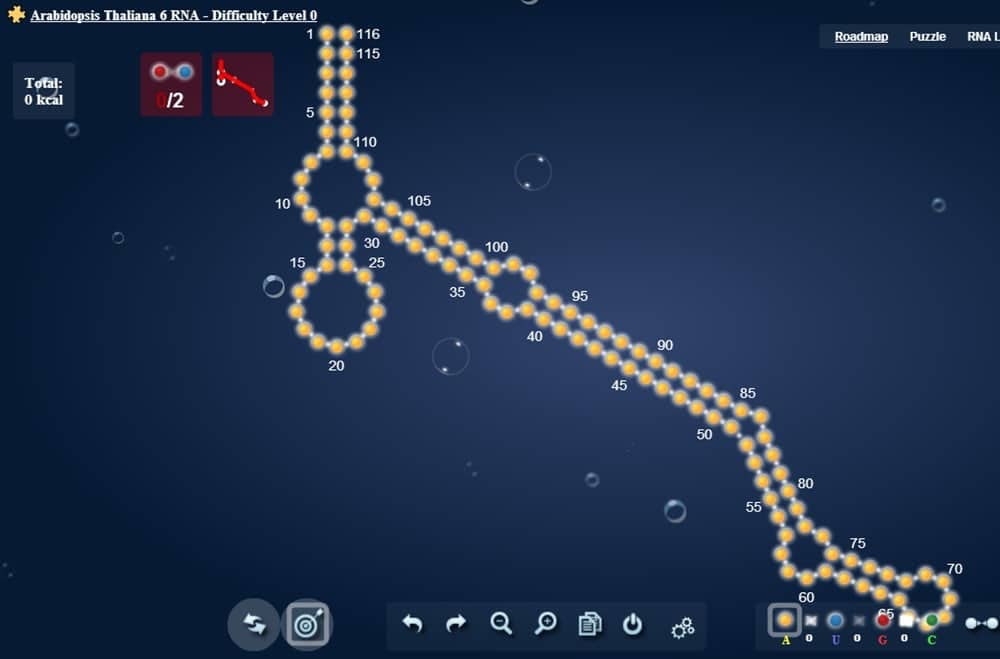
This lofty goal didn’t take long to become a reality. The game’s complexity grew alongside the molecule-making skills of the general player population. Around 100,000 registered players were active at the game’s popularity peak. In 2016, the Journal of Molecular Biology published a paper co-authored by the game players themselves. In this paper, the game players established a set of rules to determine the design difficulty of certain RNA molecular structures.
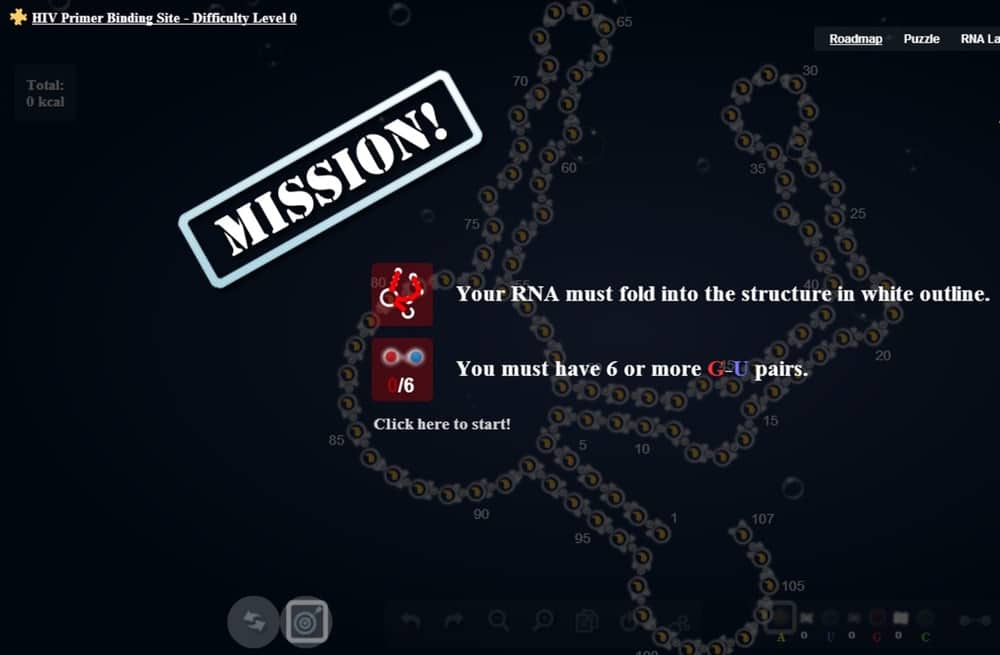
Prior to this paper, EteRNA players were called upon to design a novel molecule that would help with the creation of an accurate and simple tuberculosis blood test. Similar to that challenge, researchers now want players to virtually design a molecular on/off switch for CRISPR/Cas9. This CRISPR control switch would essentially make new research and therapies viable.
A Crisper Perspective of CRISPR
CRISPR/Cas9 is a bacterial gene-editing tool system. In 2012, researchers learned how to deploy the system (CRISPR itself, the Cas9 protein, and a “guide” RNA) in a more controlled manner. Basically, E. coli, a bacterium species residing in your gut, has the ability to read and damage viral DNA. The RNA molecules of this bacterium can use the controlled viral DNA in CRISPR as a reference to find a viral infection. The Cas9 enzyme then cuts the found viral DNA, allowing for editing.
Essentially, researchers could now recognize, cut, and replace specified DNA sequences. CRISPR is extremely powerful and has already been applied to numerous projects; viruses, human stem cells, and even corn are a few. But because it is so powerful, it’s not exactly something you want to have on and active all the time. Besides solving this problem, an “on/off” switch could also act as a timer that allows CRISPR to produce certain molecules in a scheduled way. This would be similar to how people consume medical drugs.
Right now, similar functions don’t allow researchers to know if CRISPR is completely on or off. It’s not hard to imagine how this could be problematic when you’re measuring things by the molecules.
“A switch on the CRISPR/Cas9 system can control how the genes are being expressed — that is, when and where.”
– William Greenleaf, PhD, Associate Professor of Genetics at Stanford University

$15 million of funding for this new challenge came from Stanford’s Center for Personal Dynamic Regulomes. The center’s director, Howard Chang, MD, PhD, explains the prerequisites for EteRNA: “All you need is a good internet connection, the interest, and the time.” There’s no need for a fancy computer here.
Power in Numbers
Chang discusses the philosophy behind EteRNA: “Great ideas can come from anywhere, so this is also an experiment in the democratization of science. A lot of people have hidden talents that they don’t even know about. This could be their calling.”
“Maybe there’s somebody out there who is a security guard and a fantastic RNA biochemist, and they don’t even know it.”
– Howard Chang
“There is a misconception of science as something that happens in an ivory tower by someone in a white coat with a long beard. And they are saying things and drawing things that nobody understands,” Chang elaborates. “But it’s not like that! It’s really like a puzzle that anybody can get engaged with.”
The more submitted solutions, the better, as Rhiju Das, principle investigator for EteRNA, puts it: “We’re not sure yet if there will be unforeseen problems with the Cas9 protein experimentally. That’s partially why we want as many diverse solutions as possible.”

Greenleaf and his lab will test first submissions to return the data to players in hopes of improving the design. Das elaborates on just how many solutions they’d like to see: “We’re hoping for 10,000 to 100,000 players to contribute 10 solutions each.” If this has you excited to play with genetics, head on over to EteRNA now.
Looking to update your gaming rig? Check out our reviews for the Elgato Cam Link, the Mira Prism AR Headset, and the Razor Blade Pro V2.
Sources: Stanford Medicine, New Atlas





